As vispy uses OpenGL and your graphics card, panning and zooming are highly performant. Under the hood, the canvas is a object which has built-in support for features such as zooming and panning. The canvas is in the center of the viewer and contains the visual display of the data passed to napari, including Images, Points, Shapes, and other supported data types. Help contains the citation and about information. Plugins allows you to install and manage plugins and displays a list of plugins that are currently installed. Window allows you to open the integrated console, display the layer controls and layer list. View allows you to toggle full screen, the menu bar, play, display axes, the scale bar, tooltips, and the activity dock. Note: In macOS, Preferences is under the napari menu. To learn more about the Preferences menu, there is a tutorial designed for developers here. Preferences allows you to personalize napari to some degree. The main menu consists of the File, View, Window, Plugins, and Help options.įile has the options to open files, folders, and samples, save layers and screenshots,copy screenshots to clipboard and, in the Windows version, preferences.Īll the options on the File menu are relatively self-explanatory except Preferences on the Windows version of napari. We’ll go through each of these in the next sections. The image below has the areas of the viewer labeled: The viewer is organized into a few key areas which are explained in the next sections: Now we will continue exploring the rest of the viewer. For more information on adding images to the viewer see the image layer guide. Imshow and the add_image methods accept any numpy-array like object as input, including n-dimensional arrays. The fastest way to open a viewer with an image on the canvas is using imshow: Let’s get started by launching a viewer with a simple 2D image. You can use this console to programmatically interact with an open viewer using the API methods illustrated in this tutorial. You can open it with the IPython console button (far left viewer button) or with the menu option Window > console. Note: There is also an IPython console available in napari, when napari is launched from the terminal, from a Python script, or when you use the napari bundled app. Open an interactive viewer) and run them.  
We will use the syntax for running the code inside a jupyter notebook with each code block below pasted into its own cell, but if you’d like to use a python script instead, simply copy and paste the code blocks into scripts with n() as the final line (this starts an event loop which will All four methods launch the same viewer, and anything related to interacting with the viewer on the screen applies equally to all of them. 
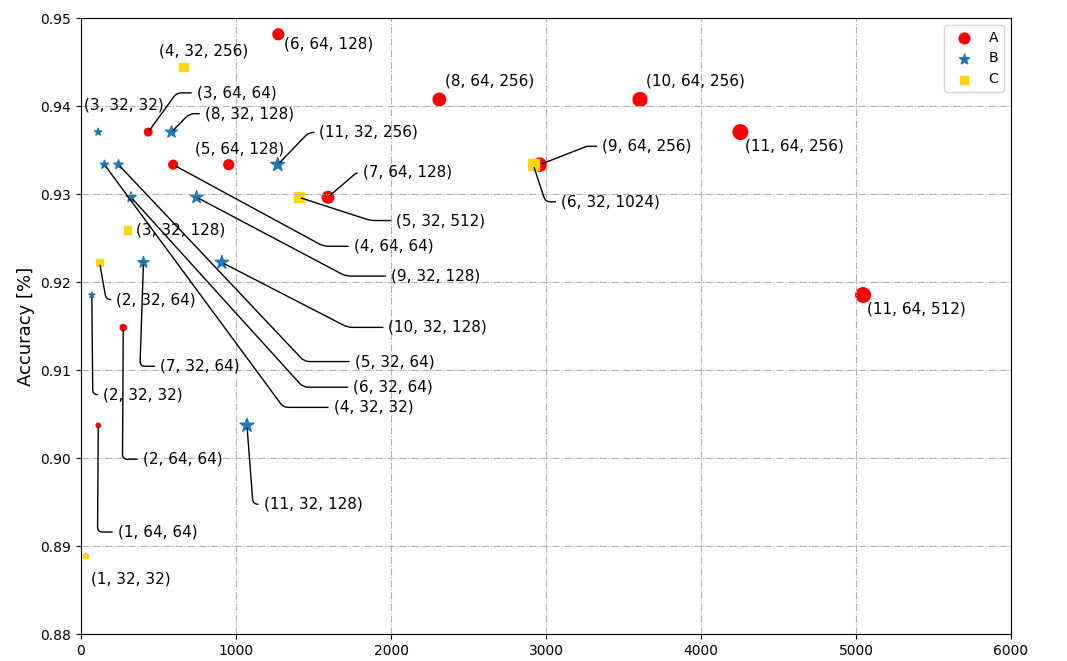
Launching the viewer ¶Īs discussed in the getting started tutorial, the napari viewer can be launched from the command-line, a python script, an IPython console, or a Jupyter notebook. At the end of the tutorial, you should understand both the layout of the viewer on the screen and the data inside of it. This tutorial will teach you about the napari viewer, including how to use its graphical user interface (GUI) and how the data within it is organized. For help launching the viewer see our getting started tutorial. For help with installation see our installation tutorial. This tutorial assumes you have already installed napari and know how to launch the viewer. Welcome to the tutorial on the napari viewer! Method to get napari style in magicgui based windows If anyone could find a solution to my problem, that would be great.Using Dask and napari to process & view large datasetsĪnnotating segmentation with text and bounding boxesĪn introduction to the event loop in napari There are other answers on overlapping annotations, but none of them are applicable to my one-liner annotation scheme. Now, it's not entirely unreadable, but what happens when I run it on a folder with 100 images? That folder has 13 images in it: 11 facial shots and 2 group pictures. The chart that it generates is less than readable: And although the function that I wrote to display the charts works fine: fig,axes = plt.subplots(2,3,figsize=(15,7))Īxes.scatter((zip(*xy_a)), (zip(*xy_a)))įor i, txt in enumerate(xy_na): axes.annotate(txt, ((zip(*xy_a)),(zip(*xy_a))))įor i, txt in enumerate(xy_na): axes.annotate(txt, ((zip(*xy_b)),(zip(*xy_b))))įor i, txt in enumerate(xy_na): axes.annotate(txt, ((zip(*xy_c)),(zip(*xy_c))))įor i, txt in enumerate(xy_na): axes.annotate(txt, ((zip(*xy_d)),(zip(*xy_d))))Īxes.set_xlabel('Dimentional Mean')įor i, txt in enumerate(xy_na): axes.annotate(txt, ((zip(*xy_e)),(zip(*xy_f))))Īxes.set_ylabel('Dimentional Mean')įor i, txt in enumerate(xy_na): axes.annotate(txt, ((zip(*xy_f)),(zip(*xy_f))))įig.suptitle("Folder: ".format ((str(folder_path)).split("/"))) I'm writing a script to classify images based on their features: Histogram, color balance, keypoint count, etc. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
